Problem background
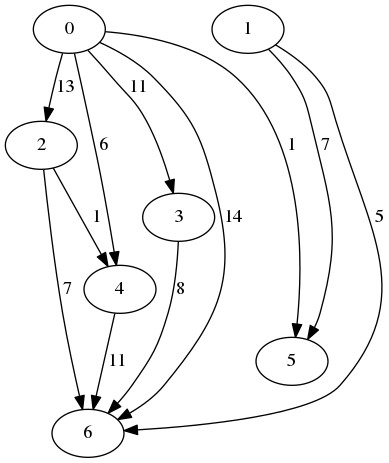
Assume that we have a large graph containing many nodes and edges. For this graph, we need to compute the number of shared neighbors for each pair of nodes.
An instance example

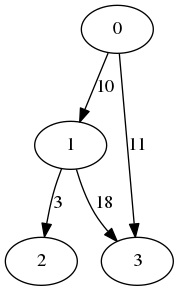
For nodes 0 and 1, the result is 1 as the nodes share a single neighbor, which is node 2. For nodes 0 and 3, the result is 2 as both of them share nodes 2 and 4 as neighbors.
Naive solution
Assume that the input graph is represented in python networkx graphs. A trivial solution is to compute for all pairs the number of shared neighbors by neighbors intersection:
from itertools import combinations
import networkx as nx
from networkx.generators.random_graphs import gnp_random_graph
g: nx.Graph = gnp_random_graph(1000, 0.2) # get a graph
shared_neighbors = {}
for node_1, node_2 in combinations(g.nodes, r=2):
shared_neighbors[(node_1, node_2)] = (
sum(1 for _ in nx.common_neighbors(g, node_1, node_2))
)
This code is relatively trivial. The most expensive part of the code is getting the common neighbors. This code can be improved by computing shared neighbors faster.
Almost naive solution
A straight forward approach to optimize the performance is by indexing the neighbors beforehand in sets for each node. Using that, we don’t need to access the graph’s data structure, and avoid an expensive iteration.
from itertools import combinations
import networkx as nx
from networkx.generators.random_graphs import gnp_random_graph
g: nx.Graph = gnp_random_graph(1000, 0.2) # get a graph
neighbors = {node: set(g.neighbors(node)) for node in g.nodes}
for node_1, node_2 in combinations(g.nodes, r=2):
shared_neighbors[(node_1, node_2)] = (
len(neighbors[node_1] & neighbors[node_2])
)
The difference between the naive solution and the almost naive one is significant. On average, an example of 1000 nodes takes 90 seconds using the naive solution, and 3 seconds using the almost naive solution.
Now, can we do any better?
Optimizing using an adjacency matrix
Firstly, let us refresh our memory regarding how graphs are represented in a matrix.
Adjacency matrix
For undirected graphs, a common representation is an adjacency matrix. Denote by n the number of nodes, and denote by A a matrix of size n×n. The cell A[i,j]=1 if the nodes i and j are adjacent, and 0 otherwise.
For example, the following graph:

Is represented by:
| 0 | 1 | 0 | 1 |
| 1 | 0 | 0 | 1 |
| 0 | 0 | 0 | 1 |
| 1 | 1 | 0 | 1 |
Simply stating, each cell corresponds to a “potential” edge. If an edge connecting the ith node to a jth node exists in the graph, the cell value is 1, otherwise it is 0.
Intersection using multiplication
Let us also remember how matrix multiplication is performed. Given matrices A and B, both of size n×n, the (i,j) cell in the result is the sum of A[i,k]•B[k,j] over all values of k from 0 to n-1. Since our values are binarized, then A[i,k]•B[k,j] is either 0 or 1, where 1 means that A[i,k]=1 and B[k,j]=1, otherwise A[i,k]=0 or B[k,j]=0. Since in adjacency matrix A[i,j] indicates if i and j are adjacent, then the sum A[i,k]•B[k,j] over k is a counter of the shared neighbors of i and j.
Using the adjacency matrix above, set i=0 and j=1: A[0,0]•B[0,1]+A[0,1]•B[1,1]+A[0,2]•B[2,1]+A[0,3]•B[3,1]=0+0+0+1=1, which corresponds to the single shared neighbor, node 3.
It is easy to see that if we multiply A by itself, each cell A[i,j] will contain the number of shared neighbors by i and j.
Now, let us look at a code that does exactly that:
A = nx.to_scipy_sparse_matrix(g)
A = (A**2).todok()
The code does three simple operations:
- Convert the graph to adjacency matrix (a csr matrix in this case)
- Multiply the adjacency matrix by itself
- Convert the result matrix to dok matrix (a dictionary of keys representation)
The first two steps perform and the expected operations described for the optimizations. The last step is performed to make the comparison to the previous versions valid, as they yielded a dictionary that allowed access in O(1) to the intersection size.
On average, an example of 1000 nodes takes 3 seconds using the almost naive solution, and 0.8 seconds using the matrix-based solution. This is more than 3 times faster, and more than 100 times faster than the naive solution.
We won’t be going into two essential details of why matrix-based operations are faster, and representation by sparse matrix. Yet, we’ll assume for now that you already possess the processor-based intuition for it.
Summary
In this post, we presented a common problem of nodes close-neighborhood intersection. We presented how easy is it to get a working solution, and how to optimize by a factor of more than a 100 using a conversion of the representation to an adjacency matrix.
In general, many problems in graphs are easily translatable to matrix problems. When a performance issue arises in the original representation, a conversion to a matrix may ease the issue significantly.